SARS-CoV-2 Spike Protein
Molecular Models in collaboration with Lee 3D, have been working with life-science researchers and scientists across the UK and beyond to bring molecular structures to life using colour 3D printing. We printed the SARS-Cov-2 spike trimer for Prof. Jason McLellan (University of Texas at Austin).
Copies of the model have been gifted to the vaccine development teams at Oxford University and Imperial College London. These models are hugely beneficial as communication aids in public outreach.
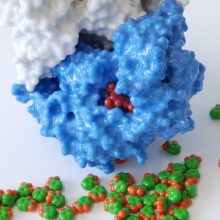
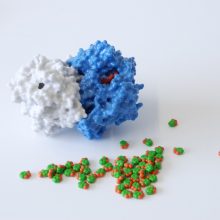
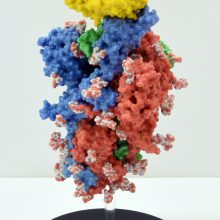
The SARS-CoV-2 spike protein is a key target for a vaccine against Covid-19. It is termed a ‘spike’ protein since dozens of copies of this molecule sit on the outside of each tiny corona virus; these give each virus a distinct spiky appearance – like a crown (or corona).
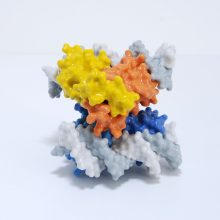
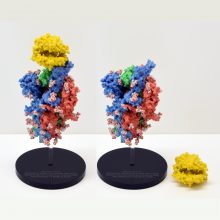
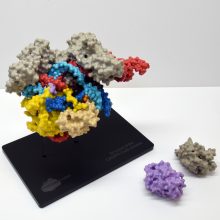
The spike protein is made of three protein chains (shown in red, green and blue on the model) that wind together to form a trimer. When a Covid-19 virus infects a human, the tip of each spike can interact with a human protein that is naturally present in the cell membrane of our respiratory epithelia – angiotensin-converting enzyme (ACE2).
The ACE2 molecule is shown in yellow and the magnets show how it can specifically attach to the spike trimer. When the spike binds to ACE2, this can lead to the virus binding and then infecting the human cell. The trimer is also covered in small sugar molecules called glycans and the biological function of these and their role in infection is still being studied.
The structure we have printed comes from cryo-EM data published by the group of Prof. Jason McLellan, University of Texas at Austin, in the journal Science in February 2020:
Wrapp et al, (2020), Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation, Science, 367, 1260–1263
Structure of the CRISPR-Cas9 complex
This model, solved by Prof. Jennifer Doudna’s team at the University of California, Berkeley, shows the crystal structure of catalytically-active Streptococcus pyogenes CRISPR-Cas9 in complex with a single-guide RNA and a double-stranded viral DNA target. The complex is primed for DNA cleavage and shows how bacteria use a CRISPR-Cas nuclease to recognise and then degrade invading viral DNA – a form of bacterial immune system. This recognition is achieved with extraordinary specificity using a guide-RNA strand that directs the Cas9 enzyme to a specific DNA sequence. This property of being able to target, cut or change any DNA sequence with exquisite precision has led to the exciting and rapidly-expanding field of CRISPR-Cas gene-editing technologies.
The colourful hands-on model we have made is ideal for use in undergraduate and postgraduate lectures and practicals, allowing students to explore, measure and appreciate the close link between molecular structure and function. The model works well in any life science degree where gene-editing is taught: from genetics to biochemistry, biotechnology to biomedical science. The model is equally useful to researchers as a visual reference and hands-on demo in science communication and school/college outreach events.
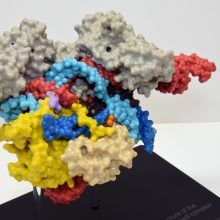
The 25 cm-long model sits on a mounting stand, from which it can be easily removed. We have used ten different colours to identify ten key features/domains of the complex. This visual key is to aid learning: the guide-RNA is in red, the DNA strands are in light and dark blue and the Cas9 enzyme domains are in seven further colours. We provide a list of each feature – and what it does – with the model. We have added optional labels (that are attached to the structure) to help students visually identify the various parts of the structure.
As the photos above show, there are three magnetically-detachable sections that help reveal where the separated DNA strands and guide-RNA are; how the nuclease active sites are positioned next to the DNA, and how Cas9 encircles the nucleic acids.
The structure we have printed comes from X-ray crystallography data (PDB code 5F9R) published by the group of Prof. Jennifer Doudna, UC-Berkeley, in the journal Science in February 2016:
Jiang et al, (2016), Structures of a CRISPR-Cas9 R-loop complex primed for DNA cleavage, Science, 351, 867–871
Structure of the F1Fo ATP Synthase
This molecular model was a collaboration with Prof. Thomas Meier at Imperial College London. It shows all twenty-six subunits that comprise a fully functioning plant ATP synthase. ATP synthases – found across all three kingdoms of life – act like molecular turbines and generate large numbers of high-energy ATP molecules for cell use. The machine operates by directing protons (H+ ions) from one side of a membrane (here, the chloroplast thylakoid membrane) to the other.
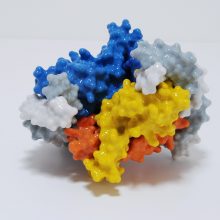
The model is coloured by subunit to aid understanding of how the motor works. It comprises two major complexes, a water soluble F1 complex and the membrane-embedded (Fo) complex. Functionally, the ATP synthase can be divided in a stationary part (called the stator) and rotary part (called the rotor). Protons enter the Fo motor by the stator a-subunit (pale blue+red), which directs them onto the c-ring rotor (wheat). On each of the 14 c-ring subunits is a glutamate residue (red spot) that conducts the protons round almost 360° to an exit point on the a-subunit. The continuous movement of this proton stream (shown in red) is converted into rotary motion. The c-ring is directly attached to a rotor shaft that comprises the epsilon (magenta), and gamma subunits (blue+yellow). So as the c-ring rotates, the attached shaft spins at the same rate.
The key feature of the model is that the Fo motor is free to rotate, providing a clear visual demonstration of how the top of the gamma subunit rotates deep within the stationary F1 head. The hexameric head is made from alternating alpha and beta subunits (dark and light green) and is held in place by the peripheral stalk, a scaffold of subunits made from the b-subunit (purple), b’-subunit (pink) and delta-subunit (orange). The ATP synthase rotation mechanism is fully reversible and can operate either in ATP synthesis mode (making ATP, using the proton-motive force), or in ATPase (hydrolysis) mode, to pump protons across the membranes and maintain the proton-motive force, depending on cell physiological conditions. The movie shows the ATP synthase operating in clockwise direction (when viewed from to of F1), which is in ATPase proton pumping direction.
The 20 cm-high model sits on a removable transparent frame, which fits into a base section. Using glossy colours to identify each of the 26 subunits and how they are positioned in relation to each other was a key part of the project brief. Also visible inside the alpha and beta headgroup are the ATP and ADP molecules, where ATP synthesis is occurring by the classic 3-stage binding-change mechanism.
The structure we have printed comes from cryo-EM data (PDB code 6FKF) and was published in the journal Science by the team of Alexander Hahn, Janet Vonck, Deryck Mills, Thomas Meier and Werner Kühlbrandt:
Hahn et al, (2018), Structure, mechanism and regulation of the chloroplast ATP synthase, Science, 360, eaat4318.
Molecular Models Gallery